Thank you, both.
Were these the models you had in mind?
library(nlme)
library(lme4)
library(emmeans)
library(ggplot2)
library(dplyr)
── 1. Data
url_raw ← “https://github.com/jorgemmteixeira/nlmeU-datset/raw/main/data/armd0.rda”
load(url(url_raw))
Filter to post-baseline rows (baseline visual0 remains as a fixed covariate)
armd0_sub ← armd0[armd0$time > 0, ]
armd0_sub$time_f_ord ← factor(armd0_sub$time.f, ordered = TRUE)
armd0_sub$subject ← as.factor(armd0_sub$subject)
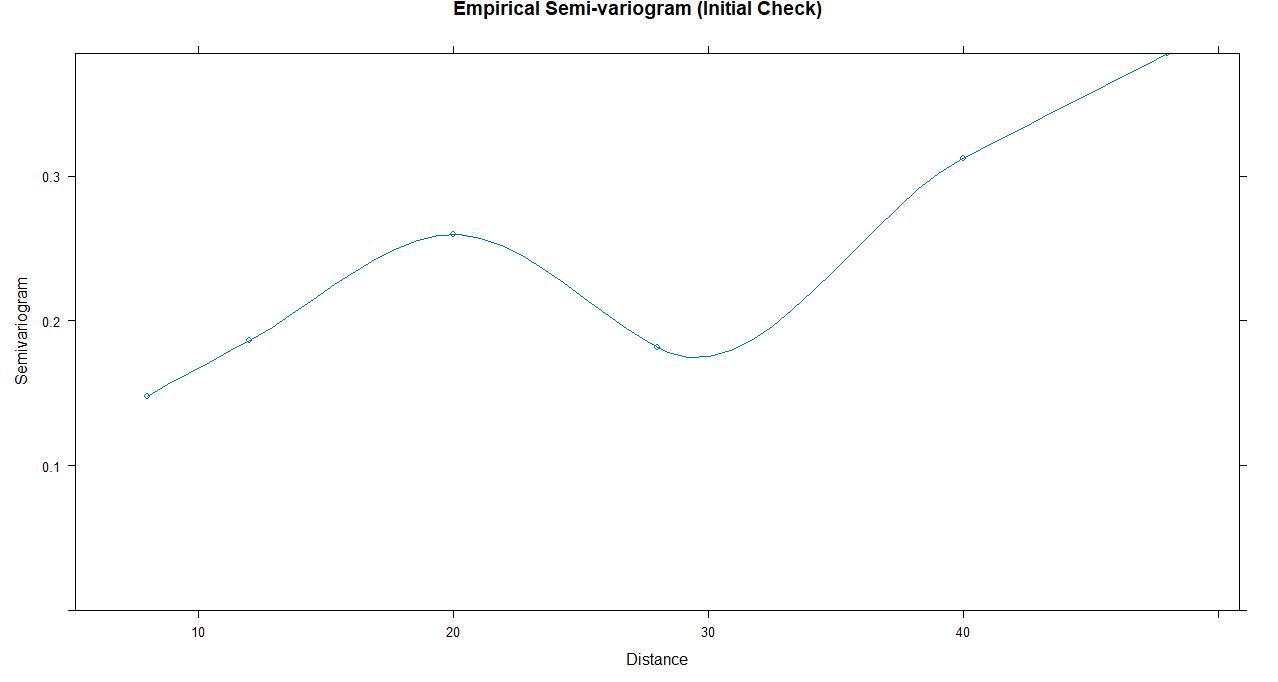
Compute and plot empirical variogram
vgm_emp ← Variogram(gls_ols, form = ~ time | subject)
plot(vgm_emp, main = “Empirical Semi-variogram (Initial Check)”)
── 2. Model Fitting (No Random Slopes) ─────────────────────────────────────
— nlme GLS (Marginal Models) —
gls_ar1 ← gls(visual ~ visual0 + time * treat.f, data = armd0_sub,
correlation = corAR1(form = ~ tp | subject), method = “REML”)
gls_exp ← gls(visual ~ visual0 + time * treat.f, data = armd0_sub,
correlation = corExp(form = ~ time | subject), method = “REML”)
gls_unstr ← gls(visual ~ visual0 + time * treat.f, data = armd0_sub,
correlation = corSymm(form = ~ tp | subject),
weights = varIdent(form = ~ 1 | time.f), method = “REML”)
— lme4 LMM (Conditional Models - Random Intercept Only) —
lme_cs ← lmer(visual ~ visual0 + time * treat.f + (1 | subject), data = armd0_sub)
lme_ar1 ← lmer(visual ~ visual0 + time * treat.f + ar1(time_f_ord + 0 | subject), data = armd0_sub)
lme_diag ← lmer(visual ~ visual0 + time * treat.f + diag(time.f + 0 | subject), data = armd0_sub)
— Markov Transition Model (OLS) —
armd_lag ← armd0_sub
group_by(subject)
mutate(visual_prev = lag(visual))
filter(!is.na(visual_prev))
markov_ols ← lm(visual ~ visual0 + visual_prev + time * treat.f, data = armd_lag)
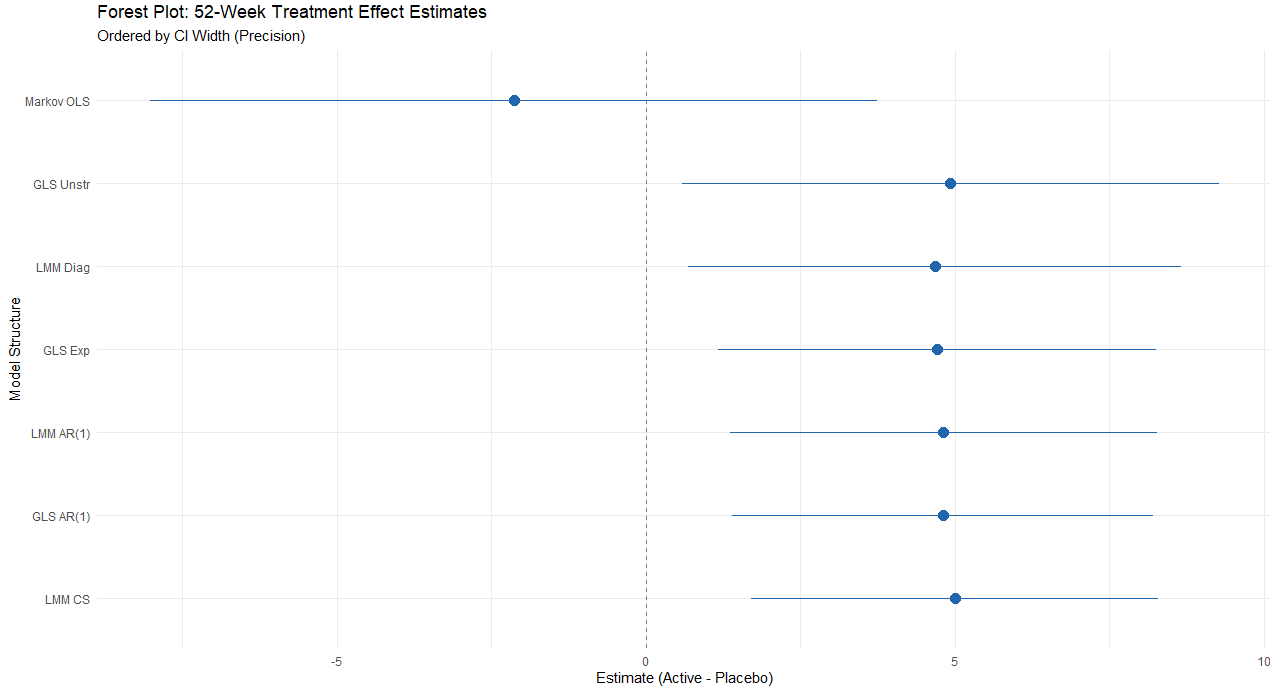
── 3. Extract Estimates and Calculate CI Widths ────────────────────────────
extract_52 ← function(mod, name) {
emm ← emmeans(mod, pairwise ~ treat.f | time, at = list(time = 52))
res ← as.data.frame(summary(emm$contrasts, infer = TRUE))
data.frame(Model = name, Estimate = res$estimate, Lower = res$lower.CL,
Upper = res$upper.CL, P_Value = res$p.value)
}
Combine all except Markov
results ← rbind(
extract_52(gls_ar1, “GLS AR(1)”),
extract_52(gls_exp, “GLS Exp”),
extract_52(gls_unstr, “GLS Unstr”),
extract_52(lme_cs, “LMM CS”),
extract_52(lme_ar1, “LMM AR(1)”),
extract_52(lme_diag, “LMM Diag”)
)
Add Markov Manually (Calculated at t=52)
m_summ ← summary(markov_ols)$coefficients
m_est ← m_summ[“treat.fActive”,1] + m_summ[“time:treat.fActive”,1] * 52
m_se ← sqrt(vcov(markov_ols)[“treat.fActive”,“treat.fActive”] +
(52^2)vcov(markov_ols)[“time:treat.fActive”,“time:treat.fActive”])
results ← rbind(results, data.frame(
Model = “Markov OLS”, Estimate = m_est, Lower = m_est - 1.96m_se,
Upper = m_est + 1.96*m_se, P_Value = 2 * (1 - pnorm(abs(m_est/m_se)))
))
── 4. Summary Table (Ordered by CI Width) ──────────────────────────────────
final_table ← results
mutate(CI_Width = Upper - Lower)
arrange(CI_Width)
select(Model, Estimate, Lower, Upper, CI_Width, P_Value)
print(final_table)