David Spiegelhalter (2019) suggested in his recent book that learning from data is a bit of a mess and that confidence intervals have baffled generations of students. The definition of a P-value being the probability of an observed value or some other more extreme hypothetical values conditional on another single hypothetical value convinces many that statistics is impossible to truly understand. However, all students understand the idea of someone else replicating or confirming their own observations in clinics and labs and the probability or expected frequency of this happening.
I show in my revised preprint (https://arxiv.org/pdf/2512.13763) that if a second study has an infinite sample size, then the probability of replicating the first study result with the same sign difference is numerically equal to one minus the one-sided P value (e.g. 1-0.025 = 0.975). If we wish the replicating study to be 1.96 SEM greater than the minimum same sign (to get a P value of 0.025 one sided again) the probability of this happening is 0.5. However, if the replicating study is the same size as the original study, then the probability of getting P ≤ 0.025 is only 0.283 and the probability of a same sign result is 0.917 as shown by Killeen (2005).
In this pre-print, I use mathematical notation instead of Excel notation. I make familiar assumptions such as a study was done perfectly with no bias etc, that unconditional prior probabilities are uniform for the scale of possible true values and possible sample values and that the probability of replication should be calculated by adding variances.
Figure 1.
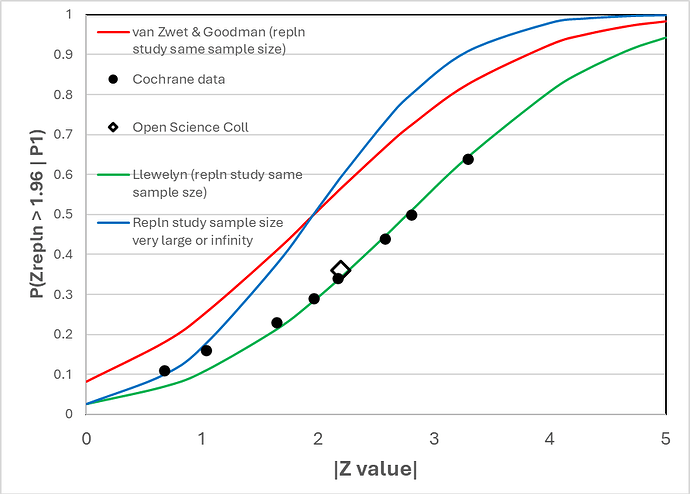
Figure 1 shows the correspondence between the frequency of replication with a P value of 0.05 two-sided (or 0.025 one-sided) estimated by van Zwet and Goodman (2022) from the Cochrane data base (represented by solid black round symbols) and various replication models (continuous lines of various colours). The result of the Open Science Collaboration is represented by the black rimmed diamond symbol.
The green line represents the expected frequency of replication using my model based on the replication study being of the same size as the original study. They correspond closely with the Cochrane and Open Science frequencies.
The red line is when the van Zwet expression (https://discourse.datamethods.org/t/some-thoughts-on-uniform-prior-probabilities-when-estimating-p-values-and-confidence-intervals/7508/310?u=huwllewelyn ) is applied to a replication study of the same sample size. It does not correspond to the Cochrane data as pointed out already by van Zwet and Goodman in their 2022 paper. Below a Z value of 1.96, when the replicating study sample size is modest and the same as in the original study (red line), the probability of replication is higher than when the replicating sample size is infinitely large (blue line), suggesting that there is a problem with their expression.
The blue line corresponds to my model when the replicating study’s sample size is infinitely large. It also corresponds to the result of applying the model and expression used by van Zwet if the sample size of the replicating study is infinitely large. Both expressions give the same result.
It appears that the assumptions made above for my expression, including an assumption of little bias, are reasonable for the RCTs in the Cochrane data base and also for the Open Science study. However, if there is a suspicion that any of the assumptions are invalid for some study, then the study result could be rejected or an attempt at correction could be made by using Bayesian methods.
An interesting situation arises with respect to the demand for two study results with P values of ≤ 0.05 two-sided or ≤ 0.025 one-sided. Figure 1 shows the probability of getting such a second significant result conditional on the same sample size and the first P value. Thus, if the first P value is 0.025 one-sided, the probability of a second study of the same size getting a significant result is only 0.283.
In order to get a 80% chance of getting a significant result again when P = 0.025, we would need a second sample size of just over 4 times the first. Similarly, if 4 separate studies of the same size were done, there is a probability of 0.28 of getting a significant result after the 1st study, 0.5 after the 2nd study, 0.67 after the 3rd study and 0.79 after the 4th study. This only applies on average before any repeat studies are done. However, if we combined results cumulatively, then the probability of replication will change as more data comes in.
References
Spiegelhalter, D. The Art of Statistics: Learning from Data. Penguin Random House, 2019.
van Zwet EW, Goodman SN. How large should the next study be? Predictive power and sample size requirements for replication studies. Statistics in Medicine. 2022; 41(16): 3090–3101. doi:10.1002/sim.9406.
Killeen PR. An alternative to null-hypothesis significance tests. Psychological Science. 2005; 16:345–353.