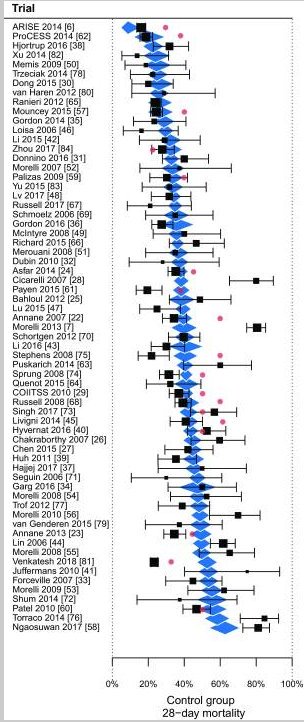
In this study, Seymour and colleagues published an extensive analysis looking to characterize clinical phenotypes for patients with Sepsis. They used several analytic techniques, and several datasets of patients to achieve this objective, and as such there is a lot to unpack. I’ll start with a few questions and thoughts I had.
-
Philosophically, do different phenotypes for the sepsis syndrome make sense?
I’d say yes. Sepsis is known to be a heterogeneous disease, with patients presenting with a wide range of symptoms and severity. The most recent consensus task force defined sepsis as infection causing a dysregulated host response, and this dysregulation they described in terms of the measurable organ dysfunction. However, within each of these components you can see where the heterogeneity arises. For infection, there are numerous types of organisms (bacteria, fungi, possibly virus), species of organism, and locations in the body (e.g. urinary tract, lungs, skin) where they may infect causing different pathophysiologic changes. For organ dysfunction, which organ is affected can have a very different physiologic impact that may aggravate the disease (e.g. gastrointestinal disruption may increase endotoxin release) or modify treatment effectiveness (e.g. increased capillary permeability shifts Starling forces, impacting hemodynamics during resuscitation). My bottom line: phenotypes for of infection, manifestations of organ dysfunction, and different patient responses should be expected. I think the uncertainty inherent to these patients is well explored in this editorial. https://www.atsjournals.org/doi/abs/10.1513/AnnalsATS.201809-646PS -
Can phenotypes be identified statistically?
If we accept phenotypes should exist, then the question is how to identify them. I think the goal would be to identify what physiologic differences are occurring in each phenotype as this may help clinicians better understand the disease process, therefore design targeted interventions. But here is where I need help from Statisticians / Machine Learning experts - how do the approaches these authors use identify phenotypes, and what are the assumptions inherent to these approaches? The word “classification” is something I am wary of as breaking patients into groups based on characteristics common to that group may not translate to actual underlying physiologic differences in their disease that are more likely to predict treatment response. -
How should phenotypes be incorporated into future studies?
Assuming phenotypes do exist and have a biologic basis, I see two problems for studies. The first is to accurately identify which phenotype a patient belongs using available information. I wouldn’t advocate for specific criteria - as we know, anytime a threshold is used to determine abnormal from normal in a continuous measure, there is inevitably information lost. But in order to use these phenotypes in future studies, we need some sort of approach to “classify” patients without adopting this harmful practice. Perhaps there is a probabilistic approach, appropriately weighting the characteristics the authors described for each phenotype? The second problem is how should these classifications be incorporated into future statistical models. Should cohorts be restricted to only patients of one phenotype? This will reduce generalizability to marginal patients between phenotypes. Should phenotypes be entered as covariates to adjust for their baseline risk for the outcome? There is a risk of incorporation bias with this approach as clinical variables are likely necessary to define the phenotypes.
Looking for comments on any of the above!