I’m trying to develop a better understanding and intuition for the transition model and was wondering how you choose to model the correlation structure of yprev and time. I’m also having some concerns regarding error rates.
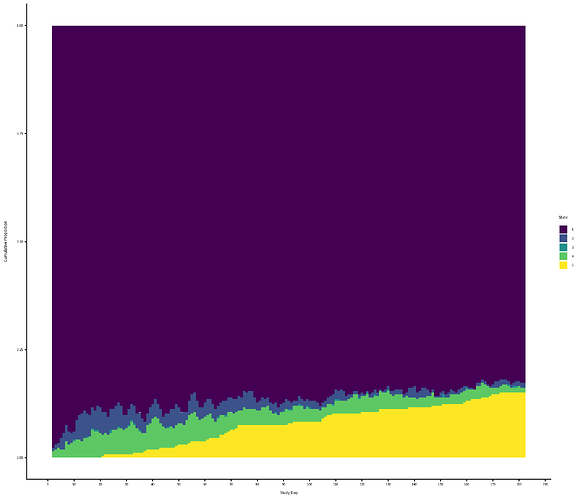
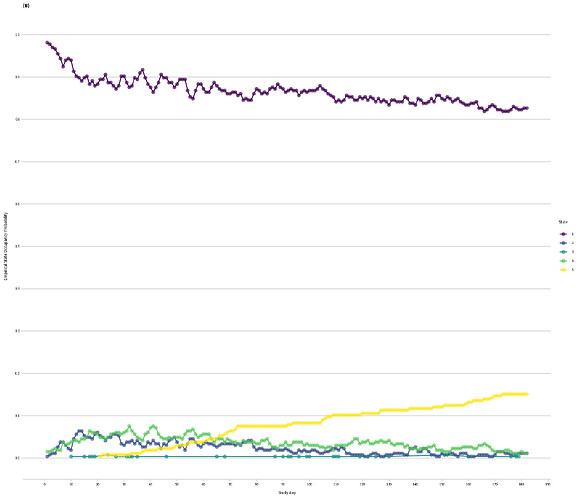
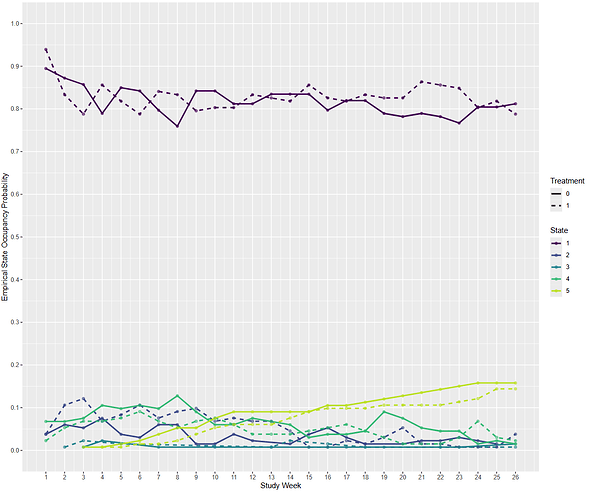
For context, I’m using a historical dataset of Stage IV cancer patients who were not in an RCT: 260 patients followed for 26 weeks. I have weekly data on whether they are at home [1], in hospital for planned treatment [2], visited the ER [3], were in hospital for an emergency [4], or dead [5]. I had to collapse state [3] and [4], because [3] is very rare. Patients spent most of their time in state 1, see empirical SOPs.
Empirical SOPs
I’ve randomly divided the patients into ‘treated’ and ‘control’ to mimic a null RCT.
Questions about modelling time and yprev
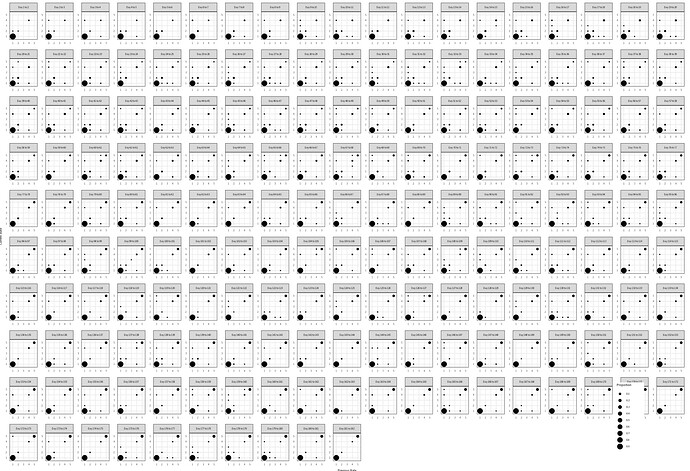
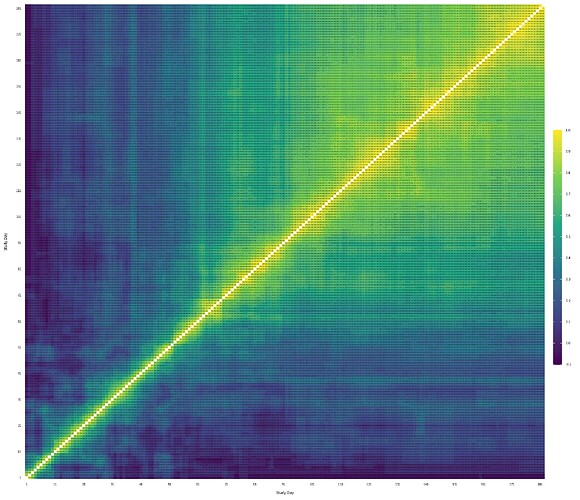
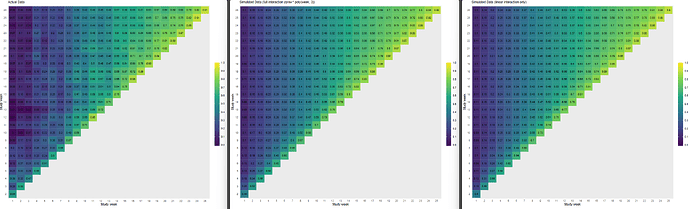
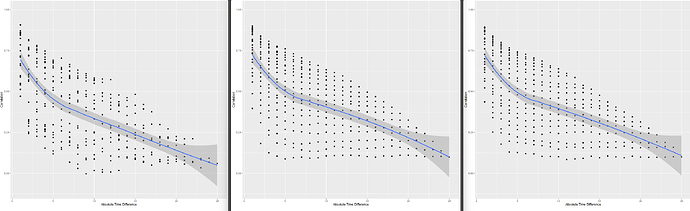
Following your paper, and chapter 22 from RMS, I constructed the empirical correlation matrix and variogram and compared those to plots made from simulated data, based on three different models fit to the original data: 1) no interaction between y and yprev, 2) linear interaction between y and yprev, 3) and full interaction between y and yprev. (No interaction not shown on plots, slightly worse fit than linear interaction)
Models used for simulated data
# I modelled time with a 3rd degree polynomial instead of splines because I found it easier to reconstruct the linear predictor for the simordmarkov function and took the median of the parameter estimates instead of all draws.
model_full_int <-
blrm(
formula = y ~ tx + pol(week, 3) * yprev + ecog_fstcnt + diagnosis + age + gender + albumin + c_reactive_protein,
data = df,
ppo = ~ week,
cppo = function(y) y,
iter = 2000,
chains = 4
)
model_lin_int <-
blrm(
formula = y ~ tx + pol(week, 3) + yprev + yprev %ia% week + ecog_fstcnt + diagnosis + age + gender + albumin + c_reactive_protein,
data = df,
ppo = ~ week,
cppo = function(y) y,
iter = 2000,
chains = 4
)
model_no_int <-
blrm(
formula = y ~ tx + pol(week, 3) + yprev + ecog_fstcnt + diagnosis + age + gender + albumin + c_reactive_protein,
data = df,
ppo = ~ week,
cppo = function(y) y,
iter = 2000,
chains = 4
)
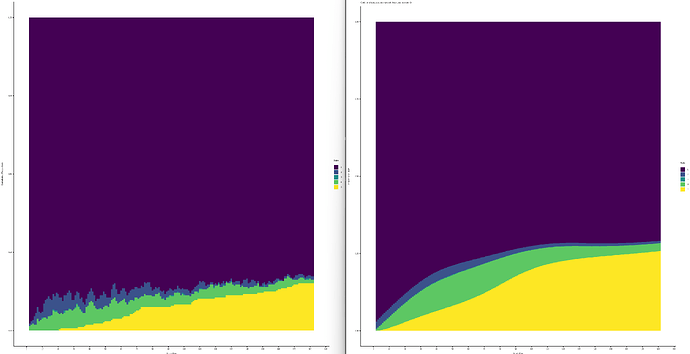
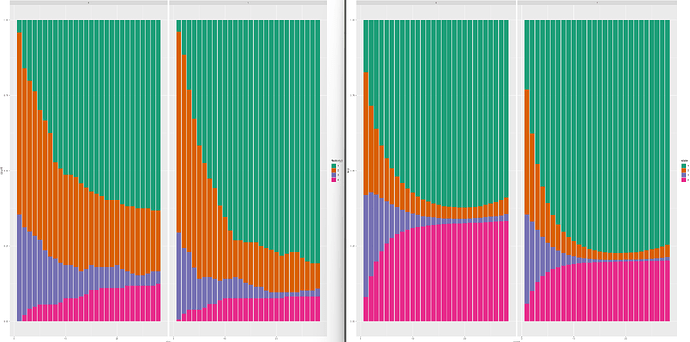
Empirical | Full Interaction | Linear Interaction
Mean absolute difference between empirical and model based correlation coefficients as a function of week:
| model |
1 |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
11 |
12 |
13 |
14 |
15 |
16 |
17 |
18 |
19 |
20 |
21 |
22 |
23 |
24 |
25 |
26 |
| full_int |
0.05 |
0.06 |
0.08 |
0.06 |
0.09 |
0.07 |
0.12 |
0.08 |
0.04 |
0.06 |
0.03 |
0.06 |
0.09 |
0.05 |
0.05 |
0.06 |
0.05 |
0.05 |
0.05 |
0.06 |
0.05 |
0.05 |
0.05 |
0.07 |
0.05 |
0.05 |
| lin_int |
0.06 |
0.06 |
0.09 |
0.06 |
0.10 |
0.08 |
0.14 |
0.10 |
0.05 |
0.08 |
0.04 |
0.06 |
0.10 |
0.06 |
0.05 |
0.06 |
0.05 |
0.05 |
0.05 |
0.07 |
0.06 |
0.06 |
0.05 |
0.07 |
0.06 |
0.06 |
| no_int |
0.06 |
0.06 |
0.11 |
0.10 |
0.14 |
0.12 |
0.17 |
0.12 |
0.07 |
0.10 |
0.06 |
0.08 |
0.11 |
0.07 |
0.05 |
0.06 |
0.05 |
0.05 |
0.06 |
0.07 |
0.06 |
0.07 |
0.06 |
0.08 |
0.07 |
0.07 |
- When I compare the empirical correlation matrix and variogram to the simulated ones, none of them look particularly great compared to the fit of the simulated data for the VIOLET trial.
- I lack experience, so I’m wondering if this approximates the empirical structure well enough? Is there any guidance on what’s not good enough? The full interaction seems to be closest to the correlation structure, but I’d be tempted to use the linear interaction only, for parsimony.
- I’m also a bit confused if not getting the correlation structure approx. right affects the type I assertion rate? I would assume it does if I choose an estimand based on SOPs and not just the beta for treatment, like time at home?
Type I Assertion Rate Concerns
I ran 1200 simulations where I randomly divided the dataset into ‘control’ and ‘intervention’ and fitted the following three frequentist models:`
m1 <- vglm(
ordered(y) ~ tx + rcs(time, 4) + yprev,
cumulative(reverse = TRUE, parallel = FALSE ~ rcs(time, 4)),
data = data)
m2 <- vglm(
ordered(y) ~ tx + rcs(time, 4) + yprev + time %ia% yprev,
cumulative(reverse = TRUE, parallel = FALSE ~ rcs(time, 4)),
data = data)
m3 <- vglm(
ordered(y) ~ tx + rcs(time, 4) + yprev + ypprev,
cumulative(reverse = TRUE, parallel = FALSE ~ rcs(time, 4)),
data = data_yppre
The type I error rate is > 0.05 when choosing alpha of 0.05 as a cut-off:
| Simulation |
Type I Error (MCSE) |
| Non-linear |
0.068 (0.008) |
| Linear |
0.068 (0.008) |
| Second-order |
0.072 (0.008) |
I also ran a simulation using bootstrapping whole patients and increasing the sample size to 500, and the error/assertion rate was even higher, ~ 10%.
I honestly don’t understand why I’m seeing this error inflation. This seems like a problem, but maybe my approach is flawed? Why could I be observing error rates > 0.05?
When using blrm() with default priors, linear constraint on ppo for time, and P(any benefit) > 0.95 for making an assertion of benefit, the error rates are very similar compared to the frequentist models:
| Model |
Point Estimate |
Rejection Rate |
Bias |
Coverage |
Empirical SE |
Model SE |
| no_int |
0.002 (0.003) |
0.069 (0.007) |
0.002 (0.003) |
0.929 (0.007) |
0.105 (0.012) |
0.095 (0.000) |
| linear_int |
0.002 (0.003) |
0.072 (0.007) |
0.002 (0.003) |
0.932 (0.007) |
0.105 (0.013) |
0.095 (0.000) |
| full_int |
0.002 (0.003) |
0.071 (0.007) |
0.002 (0.003) |
0.932 (0.007) |
0.106 (0.013) |
0.095 (0.000) |
| no_int_cluster |
0.001 (0.004) |
0.041 (0.006) |
0.001 (0.004) |
0.957 (0.006) |
0.128 (0.013) |
0.133 (0.000) |
The only thing that made a difference, was adding + cluster(id), wich reduced the error / assertion rate. The mean SD of the random effect was about 0.6 across the simulations. Does this mean that the increased error rates are because the first-order (and second-order) markov assumptions do not hold, and the random effect accounts for that?
Sorry for the long post. Any insights would be very much appreciated.
Edit:
To add to my confusion:
When taking simulated data (dataset of 200k patients) without an interaction between time and yprev, i.e., based on this model:
# states three and four were already collapsed during model fitting
vm <- vglm(
ordered(y) ~ tx + week + I(week^2) + I(week^3) + yprev + ecog_fstcnt + diagnosis,
cumulative(reverse = TRUE, parallel = FALSE ~ week + I(week^2) + week + I(week^3)),
data = df
)
And then running 1000 simulation on this data with n = 250 and using this vglm model, which correctly models time and yprev:
m <- vglm(
ordered(y) ~ tx + poly(time, 3) + yprev,
cumulative(reverse = TRUE, parallel = FALSE ~ poly(time, 3)),
data = data)
I still get inflated Type I error rates of .068 (0.008).
When I increase the sample size per iteration to 1000, the error rate goes down to 0.055 (0.007). Why would it be a matter of sample size? If I had left in state 3 in the simulated data, that might have created problems, because it was so rare, but I already collapsed states 3 and 4 before fitting the model for the simulation, so that can’t be the problem.