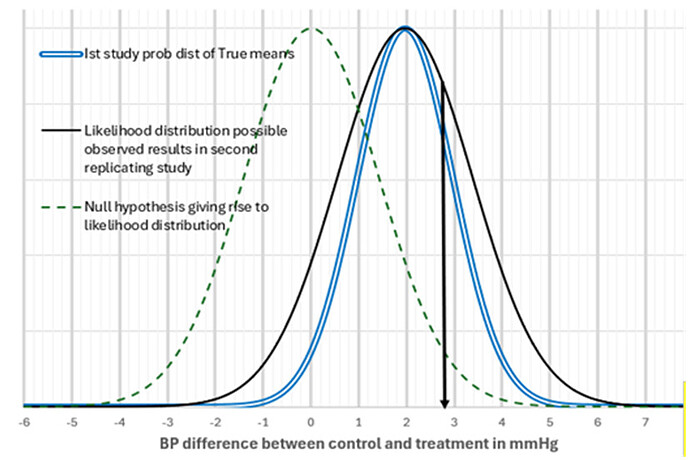
I’m surprised the OPs method gives numerically similar results as the method from my 2022 paper with Steve Goodman without using our data.
Apologies for the following long post. I hope at least it’s educational. To make the post self-contained, I’ll re-iterate my notation. Let beta denote the (unobserved) true effect effect of the treatment and let b be a normally distributed estimator with mean beta (i.e. b is unbiased) and (known) standard error s. Define the z-statistic z=b/s and the signal-to-noise ratio SNR=beta/s. The z-statistic has the normal distribution with mean SNR and standard deviation 1. In other words, z is the sum of SNR and an independent standard normal error. Let z_repl be the sum of the same SNR, but another independent standard normal error.
We’re interested in the probability of “successful replication” after having observed (or “conditional on”) the z-statistic of the original study. To be more specific, we want to evaluate
P(z_repl > 1.96 | z).
This probability depends on the distribution of the SNR. Let’s assume that the SNR has the normal distribution with mean zero and standard deviation sigma. Well known theory about bivariate normal distributions tells us that the conditional distribution of the SNR given z is again normal with mean z*sigma^2/(sigma^2 + 1) and variance sigma^2/(sigma^2 + 1).
It follows that the conditional distribution of z_repl given z is also normal with mean z*sigma^2/(sigma^2 + 1) and variance sigma^2/(sigma^2 + 1) + 1.
If I understand correctly, the OP assumed the uniform distribution for the SNR. That’s like a normal distribution with a very large sigma. In that case, the factor sigma^2/(sigma^2 + 1) is essentially 1. So, the conditional distribution of z_repl given z is normal with mean z and variance 1+1=2. For various values of z, the conditional probability that z_repl exceeds 1.96 is
z=c(0.67,1.04,1.64,1.96,2.17,2.58,2.81,3.29)
1- pnorm(1.96,z,sqrt(2))
0.18 0.26 0.41 0.50 0.56 0.67 0.73 0.83
In my paper with Steve Goodman, we used a distribution for the SNR that we estimated from a large collection of clinical trials. It’s not a normal distribution, but it can be reasonably approximated by a normal distribution with mean 0 and standard deviation sigma=1.5. The conditional probability that z_repl exceeds 1.96 becomes
m=z*1.5^2/(1.5^2+1)
v=1.5^2/(1.5^2+1) + 1
1- pnorm(1.96,m,sqrt(v))
0.13 0.17 0.26 0.32 0.36 0.45 0.50 0.60
Now let’s turn to the OPs method. He uses the uniform distribution for the SNR, but gets similar results to using N(0,1.5). How is that possible? Well, he seems to define replication success differently, namely
P(z_repl > 2.77 | z)
It is very unusual to define one-sided significance at level 0.025 in this way. Using this definition together with the uniform prior on the SNR, we get
1 - pnorm(2.77,z,sqrt(2))
0.07 0.11 0.21 0.28 0.34 0.45 0.51 0.64
However, this is the same as
1 - pnorm(1.96,z/sqrt(2),1)
0.07 0.11 0.21 0.28 0.34 0.45 0.51 0.64
At least we’re talking about the probability of exceeding 1.96 again! Now, 1/sqrt(2) = 0.71 happens to be very close to the shrinkage factor of 1.5^2/(1.5^2+1) = 0.70 which Goodman and I used. Morover, the conditional standard deviation which we used is also not too different from 1. I believe this explains the coincidental numerical agreement between our results.